# A tutorial: k-mer-based analysis of metagenomes with sourmash
C. Titus Brown, Taylor Reiter, and Tessa Pierce
July 27, 2022
MBL STAMPS
[](https://hackmd.io/vYaK2UngTWSkKmcpQMP6NA)
[(Permanent link on GitHub)](https://github.com/mblstamps/stamps2022/blob/main/kmers_and_sourmash/README.md)
This tutorial has now been added to the sourmash docs! See [here](https://sourmash.readthedocs.io/en/latest/tutorial-lemonade.html).
---
[toc]
In this tutorial, we'll use a tool called sourmash to analyze the composition of a metagenome, both genomically and taxonomically. We'll also classify our MAGs from Monday's assembly-and-binning lecture, and integrate them into our analysis.
We'll be working entirely at the shell prompt, because the shell is excellent at dealing with long running processes and large data sets.
## Install sourmash
First, we need to install the software! We'll use conda/mamba to do this.
The below command installs [sourmash](https://sourmash.readthedocs.io/) and [GNU parallel](https://www.gnu.org/software/parallel/).
First run:
```
# leave whatever environment you're in
conda activate base
# now create a new environment
mamba create -n smash -y -c conda-forge -c bioconda sourmash parallel
```
to install the software, and then run
```
conda activate smash
```
to activate the conda environment so you can run the software.
::::info
Victory conditions: your prompt should now start with
`(smash) stamps2022@149.165.154.81:~$ `
and you should now be able to run `sourmash` and have it output usage information!!
::::
### What is sourmash? Why sourmash?
sourmash (https://sourmash.readthedocs.io/en/latest/) is software that enables k-mer Jaccard similarity and containment operations on really large data sets in pretty small amounts of memory. It is developed (mostly) by the [Lab for Data Intensive Biology](http://ivory.idyll.org/lab/) - local reps are Titus, Tessa, Taylor, and Bry :).
There are other tools that will let you do similar things but we are going to show you sourmash for three reasons:
1. we are really familiar with it and like it!
2. it is pretty robust and well-rounded software that does exactly the things we want to show you (ok, this is not a coincidence)
3. it's easy to install and runs quickly enough to do everything during one session!
## Create a working subdirectory
::::info
`conda` environments are software environments, and are very different from _directories_.
Directories contain files. They are distinct from environments: you can use one conda environment and work in many directories, or be in a single directory and switch between multiple conda environments, or mix and match as needed.
::::
Make a directory named `kmers` and change into it.
```
mkdir ~/kmers
cd ~/kmers
```
::::warning
If you log out or get disconnected in the middle of the tutorial, you'll need to run the following two commands to pick up again:
```
cd ~/kmers
conda activate smash
```
::::
## Download a database and a taxonomy spreadsheet.
We're going to start by doing a reference-based _compositional analysis_ of the lemonade metagenome from [Taylor's tutorial on assembly and binning](https://github.com/mblstamps/stamps2022/blob/main/assembly_and_binning/tutorial_assembly_and_binning.md).
For this purpose, we're going to need a database of known genomes. We'll use the GTDB genomic representatives database, containing ~65,000 genomes, for today - that's because it's smaller than the full GTDB database (~320,000) or Genbank (~1.3m), and hence faster. But you can download and use those on your own, if you like!
You can find the link to a prepared GTDB RS207 database for k=31 on the [the sourmash prepared databases page](https://sourmash.readthedocs.io/en/latest/databases.html). Let's download it to the current directory:
```
curl -JLO https://osf.io/3a6gn/download
```
This will create a 1.7 GB file:
```
ls -lh gtdb-rs207.genomic-reps.dna.k31.zip
```
::::warning
If it downloaded to `download`, you can run:
```
mv download gtdb-rs207.genomic-reps.dna.k31.zip
```
to rename it
::::
and you can examine the contents with sourmash `sig summarize`:
```
sourmash sig summarize gtdb-rs207.genomic-reps.dna.k31.zip
```
which will show you:
```
>path filetype: ZipFileLinearIndex
>location: /home/stamps2022/kmers/gtdb-rs207.genomic-reps.dna.k31.zip
>is database? yes
>has manifest? yes
>num signatures: 65703
>** examining manifest...
>total hashes: 212454591
>summary of sketches:
> 65703 sketches with DNA, k=31, scaled=1000, abund 212454591 total hashes
```
There's a lot of things to digest in this output but the two main ones are:
* there are 65,703 genome sketches in this database, for a k-mer size of 31
* this database represents 212 *billion* k-mers (multiply number of hashes by the scaled number)
If you want to read more about what, exactly, sourmash is doing, please see [Lightweight compositional analysis of metagenomes with FracMinHash and minimum metagenome covers](https://www.biorxiv.org/content/10.1101/2022.01.11.475838v2), Irber et al., 2022.
We also want to download the accompanying taxonomy spreadsheet:
```
curl -JLO https://osf.io/v3zmg/download
```
and uncompress it:
```
gunzip gtdb-rs207.taxonomy.csv.gz
```
This spreadsheet contains information connecting Genbank genome identifiers to the GTDB taxonomy - take a look:
```
head -2 gtdb-rs207.taxonomy.csv
```
will show you:
```
>ident,superkingdom,phylum,class,order,family,genus,species
>GCF_000566285.1,d__Bacteria,p__Proteobacteria,c__Gammaproteobacteria,o__Enterobacterales,f__Enterobacteriaceae,g__Escherichia,s__Escherichia coli
```
Let's index the taxonomy database using SQLite, for faster access later on:
```
sourmash tax prepare -t gtdb-rs207.taxonomy.csv \
-o gtdb-rs207.taxonomy.sqldb -F sql
```
This creates a file `gtdb-rs207.taxonomy.sqldb` that contains all the information in the CSV file, but which is faster to use than the CSV file.
## Download and prepare sample reads
Next, let's download one of the metagenomes from [the assembly and binning tutorial](https://github.com/mblstamps/stamps2022/blob/main/assembly_and_binning/tutorial_assembly_and_binning.md#retrieving-the-data).
We'll use sample SRR8859675 for today, and you can view sample info [here](https://www.ebi.ac.uk/ena/browser/view/SRR8859675?show=reads) on the ENA.
To download the metagenome from the ENA, run:
```
wget ftp://ftp.sra.ebi.ac.uk/vol1/fastq/SRR885/005/SRR8859675/SRR8859675_1.fastq.gz
wget ftp://ftp.sra.ebi.ac.uk/vol1/fastq/SRR885/005/SRR8859675/SRR8859675_2.fastq.gz
```
Now we're going to prepare the metagenome for use with sourmash by converting it into a _signature files_ containing _sketches_ of the k-mers in the metagenome. This is the step that "shreds" all of the reads into k-mers of size 31, and then does further data reduction by [sketching](https://en.wikipedia.org/wiki/Streaming_algorithm) the resulting k-mers.
To build a signature file, we run `sourmash sketch dna` like so:
```
sourmash sketch dna -p k=31,abund SRR8859675*.gz \
-o SRR8859675.sig.gz --name SRR8859675
```
Here we're telling sourmash to sketch at k=31, and to track k-mer multiplicity (with 'abund'). We sketch _both_ metagenome files together into a single signature named `SRR8859675` and stored in the file `SRR8859675.sig.gz`.
When we run this, we should see:
>calculated 1 signature for 3452142 sequences taken from 2 files
so that's how many reads there are in these two files!
If you look at the resulting files,
```
ls -lh SRR8859675*
```
you'll see that the signature file is _much_ smaller (2.5mb) than the metagenome files (~600mb). This is because of the way sourmash uses a reduced representation of the data, and it's what makes sourmash fast. Please see the paper above for more info!
Also note that the GTDB prepared database we downloaded above was built using the same `sourmash sketch dna` command, but applied to 65,000 genomes and stored in a zip file.
## Find matching genomes with `sourmash gather`
At last, we have the ingredients we need to analyze the metagenome against GTDB!
* the software is installed
* the GTDB database is downloaded
* the metagenome is downloaded and sketched
Now, we'll run the [sourmash gather](https://sourmash.readthedocs.io/en/latest/command-line.html#sourmash-gather-find-metagenome-members) command to find matching genomes.
Run gather - this will take ~6 minutes:
```
sourmash gather SRR8859675.sig.gz gtdb-rs207.genomic-reps.dna.k31.zip --save-matches matches.zip
```
Here we are saving the matching genome sketches to `matches.zip` so we can rerun the analysis if we like.
::::warning
What takes so long?
Here, sourmash is looking for an estimated overlap of 100,000 k-mers (or more) between the metagenome and each of 65,000 genomes. It just takes a hot minute, is all.
(Yes, we can make it much faster, and yes, we're working on it. But 6 minutes isn't really _that_ long, is it?)
::::
The results will look like this:
```
overlap p_query p_match avg_abund
--------- ------- ------- ---------
2.0 Mbp 0.4% 31.8% 1.3 GCF_004138165.1 Candidatus Chloroploc...
1.9 Mbp 0.5% 66.9% 2.1 GCF_900101955.1 Desulfuromonas thioph...
0.6 Mbp 0.3% 23.3% 3.2 GCA_016938795.1 Chromatiaceae bacteri...
0.6 Mbp 0.5% 27.3% 6.6 GCA_016931495.1 Chlorobiaceae bacteri...
...
found 22 matches total;
the recovered matches hit 5.3% of the abundance-weighted query
```
which we discussed a little bit in lecture. Just to revisit,
* the last column is the name of the matching GTDB genome
* the first column is the estimated overlap between the metagenome and that genome, in number of k-mers
* the second column, `p_query` is the percentage of metagenome k-mers (weighted by multiplicity) that match to the genome; this will approximate the percentage of _metagenome reads_ that will map to this genome, if you map.
* the third column, `p_match`, is the percentage of the genome k-mers that are matched by the metagenome; this will approximate the percentage of _genome bases_ that will be covered by mapped reads;
* the fourth column is the estimated abundance of this genome in the metagenome.
The other interesting number is here:
>the recovered matches hit 5.3% of the abundance-weighted query
which tells you that you should expect 5.3% of the metagenome reads to map to these 22 reference genomes.
::::success
You can try running gather without abundance weighting:
```
sourmash gather SRR8859675.sig.gz matches.zip \
--ignore-abundance
```
How does the output differ?
::::spoiler Answer(s)
The main number that changes bigly is:
>the recovered matches hit 2.4% of the query (unweighted)
which represents the proportion of _unique_ kmers in the metagenome that are not found in any genome.
This is (approximately) the following number:
* suppose you assembled the entire metagenome perfectly into perfect contigs (**note, this is impossible, although you can get close with "unitigs"**);
* and then matched all the genomes to the contigs;
* approximately 2.4% of the bases in the contigs would have genomes that match to them.
Interestingly, this is the _only_ number in this entire tutorial that is essentially impossible to estimate any way other than with k-mers.
This number is also a big underestimate of the "true" number for the metagenome - we'll explain more later :).
::::
::::warning
### GTDB subset vs entire
We're using only a small subset of GTDB here, but we could do this against all of Genbank fairly easily. It would "just" require more CPU time and more memory ;). See [sourmash gather benchmarks x genbank](https://github.com/dib-lab/2020-paper-sourmash-gather/issues/47#issue-1204856050) for some details on resource usage for searching all of Genbank microbial!
This is really what sourmash is designed for - searching ridiculously large databases. (There are no other tools that we're aware of that can handle all of genbank, although the field is evolving rapidly!)
Check out [ganon](https://academic.oup.com/bioinformatics/article/36/Supplement_1/i12/5870470), [AGAMEMNON](https://genomebiology.biomedcentral.com/articles/10.1186/s13059-022-02610-4), and [KMCP](https://www.biorxiv.org/content/10.1101/2022.03.07.482835v2.full) for some cool software that does similar things with k-mers!
::::
## Build a taxonomic summary of the metagenome
We can use these genomes to build a taxonomic summary of the metagenome using [sourmash tax metagenome](https://sourmash.readthedocs.io/en/latest/command-line.html#sourmash-tax-subcommands-for-integrating-taxonomic-information-into-gather-results) like so:
```
# rerun gather, save the results to a CSV
sourmash gather SRR8859675.sig.gz matches.zip -o SRR8859675.x.gtdb.csv
# use tax metagenome to classify the metagenome
sourmash tax metagenome -g SRR8859675.x.gtdb.csv \
-t gtdb-rs207.taxonomy.sqldb | head
```
This output shows you output at all taxonomic levels; to see just the orders, do:
```
sourmash tax metagenome -g SRR8859675.x.gtdb.csv \
-t gtdb-rs207.taxonomy.sqldb | grep order
```
this will show you all the order level classifications.
If you want to clean up the output a bit more, you can run:
```
sourmash tax metagenome -g SRR8859675.x.gtdb.csv \
-t gtdb-rs207.taxonomy.sqldb | \
cut -d, -f 2,4,7 | \
sort -t, -k 3 -rn | grep order
```
to just display the rank, taxonomic lineage, and weighted fraction of the metagenome at that rank.
At the bottom, we have a script to plot the resulting taxonomy using [metacoder](https://grunwaldlab.github.io/metacoder_documentation/) - here's what it looks like:
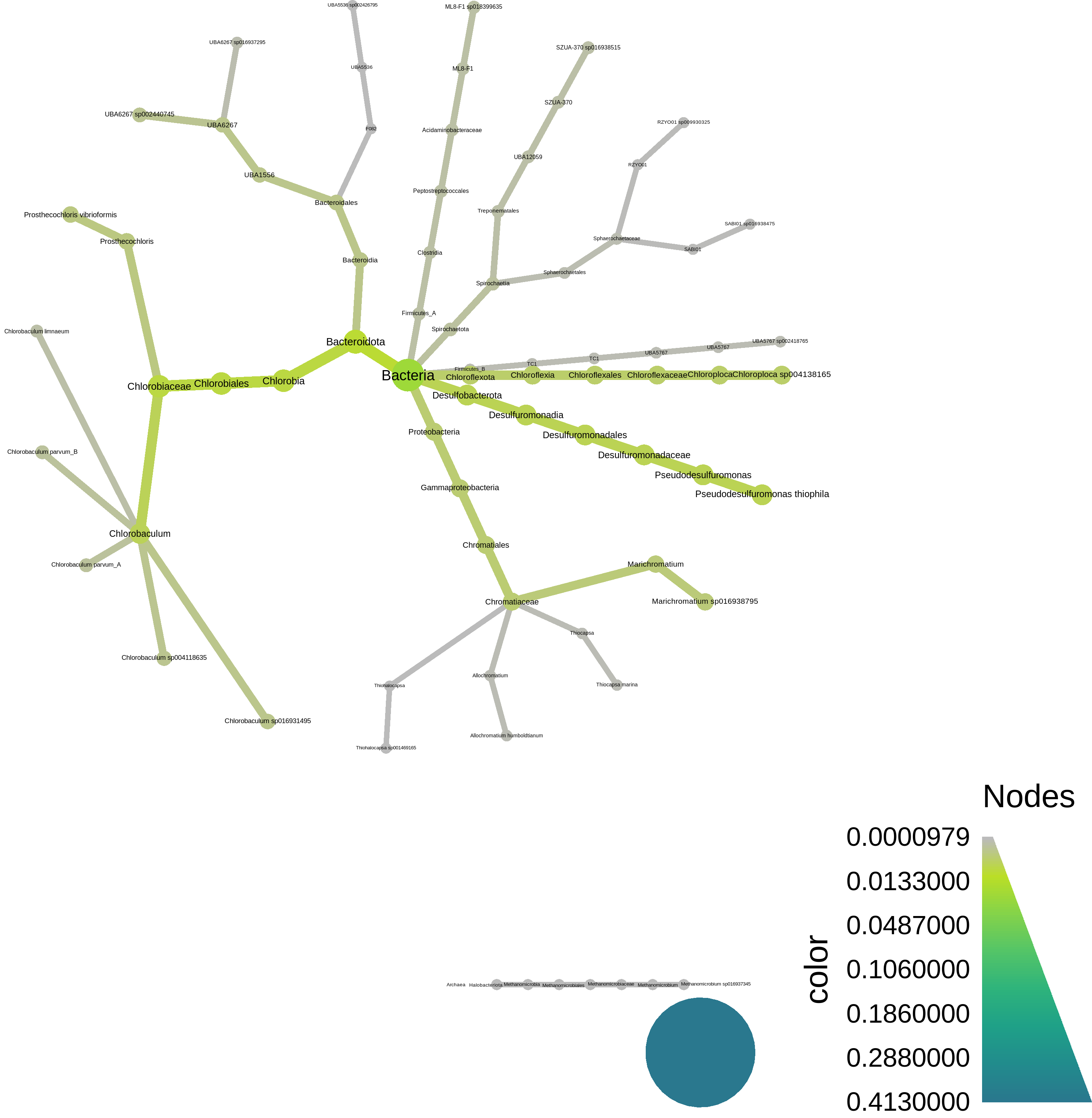
## Interlude: why reference based analyses are problematic for environmental metagenomes.
During the lecture, I showed you some preliminary data suggesting that really only a few types of metagenomes have high classification rates against Genbank. This is an environmental metagenome, and you can see that we're estimating only 5.3% of it will map to GTDB reference genomes!
Wow, that's **terrible**! Our taxonomic and/or functional analysis will be based on only 1/20th of the data!
What could we do to improve that?? There are two basic options -
(1) Use a more complete reference database, like the entire GTDB, or Genbank. This will only get you so far, unfortunately. (See exercises at end.)
(2) Assemble and bin the metagenome to produce new reference genomes!
There are other things you could think about doing here, too, but these are probably the "easiest" options. And what's super cool is that we've already _done_ the second one. So can we include that in the analysis??
Yes, yes we can! We can integrate the three MAGs that Taylor generated during her tutorial into the sourmash analysis.
We'll need to:
* download the three genomes;
* sketch them with k=31;
* re-run sourmash gather with both GTDB _and_ the MAGs.
Let's do it!!
## Update gather with information from MAGs
First, download the MAGs:
```
# Download 3 MAGs generated by ATLAS
curl -JLO https://osf.io/fejps/download
curl -JLO https://osf.io/jf65t/download
curl -JLO https://osf.io/2a4nk/download
```
This will produce three files, `MAG*.fasta`.
Now sketch them:
```
sourmash sketch dna MAG*.fasta --name-from-first
```
here, `--name-from-first` is a convenient way to give them distinguishing names based on the name of the first contig in the FASTA file; you can see the names of the signatures by doing:
```
sourmash sig describe MAG1.fasta.sig
```
Now, let's re-do the metagenome classification with the MAGs:
```
sourmash gather SRR8859675.sig.gz MAG*.sig matches.zip -o SRR8859675.x.gtdb+MAGS.csv
```
and look, we classify a lot more!
```
overlap p_query p_match avg_abund
--------- ------- ------- ---------
2.3 Mbp 12.1% 99.9% 39.4 MAG2_1
2.2 Mbp 26.5% 99.9% 92.4 MAG3_1
2.0 Mbp 0.4% 31.8% 1.3 GCF_004138165.1 Candidatus Chloroploc...
1.9 Mbp 0.5% 66.9% 2.1 GCF_900101955.1 Desulfuromonas thioph...
1.0 Mbp 2.7% 100.0% 20.3 MAG1_1
0.6 Mbp 0.3% 23.2% 3.1 GCA_016938795.1 Chromatiaceae bacteri...
0.6 Mbp 0.1% 24.5% 2.1 GCA_016931495.1 Chlorobiaceae bacteri...
...
found 24 matches total;
the recovered matches hit 43.5% of the abundance-weighted query
```
::::warning
Question 1: what "interesting" features do you note about the MAG matches?
::::spoiler Some answers:
(1) The three MAG matches are all ~100% present in the metagenome.
(2) They are all at high abundance in the metagenome, because assembly needs genomes to be ~5x or more in abundance in order to work!
(3) Because they're at high abundance and 100% present, they account for _a lot_ of the metagenome!
::::
::::warning
Question 2: we found 22 matches with the first gather, and we found 24 matches this time, using three MAGs. All three MAGs are found; shouldn't we have _25_ matches?
What is the likely reason we have 24 matches, instead of 25?
::::spoiler Answer
It turns out the MAGs partially overlap some genomes in GTDB!
```
sourmash search MAG2.fasta.sig matches.zip --threshold=0
```
::::
::::warning
Question 3: why do we not get to 100% classification of the metagenome?
::::spoiler Answers
(1) most of the constitutent genomes aren't in the reference database;
(2) not everything in the metagenome is high enough coverage to bin into MAGs;
(3) not everything in the metagenome is bacterial or archaeal, and we didn't do viral or eukaryotic binning;
(4) some of what's in the metagenome k-mers may simply be erroneous (although with abundance weighting, this is likely to be a small chunk of things)
::::
## Classify the taxonomy of the MAGs; update metagenome classification
Now we can also classify the genomes and update the taxonomic summary of the metagenome!
First, classify the genomes using GTDB; this will use trace overlaps between contigs in the MAGs and GTDB genomes to tentatively identify the _entire_ bin.
```
for i in MAG*.fasta.sig
do
# get 'MAG' prefix. => NAME
NAME=$(basename $i .fasta.sig)
# search against GTDB
echo sourmash gather $i gtdb-rs207.genomic-reps.dna.k31.zip \
--threshold-bp=5000 \
-o ${NAME}.x.gtdb.csv
done | parallel
```
(This will take about a minute.)
::::success
What are we doing with all the for loop stuff!?
::::spoiler
Well, we've got three genomes to classify, and not a lot of time to do it in! So the for loop etc above does the following:
* for each of the three MAGs,
* extract the prefix (MAG1, MAG2, MAG3),
* create a sourmash command to search them against GTDB,
* and then use [GNU parallel](https://www.gnu.org/software/parallel/) to run all three commands at the same time.
If we didn't want to be fancy and fast, we could have just typed out:
```
sourmash gather MAG1.fasta.sig \
gtdb-rs207.genomic-reps.dna.k31.zip \
--threshold-bp=5000 \
-o MAG1.x.gtdb.csv
sourmash gather MAG2.fasta.sig \
gtdb-rs207.genomic-reps.dna.k31.zip \
--threshold-bp=5000 \
-o MAG2.x.gtdb.csv
sourmash gather MAG3.fasta.sig \
gtdb-rs207.genomic-reps.dna.k31.zip \
--threshold-bp=5000 \
-o MAG3.x.gtdb.csv
```
but every time I type more than two commands like this, I make a mistake (and you will too!); and it's nice to run things fast, innit??
::::
If you scan the results quickly, you'll see that one MAG has matches in genus Prosthecochloris, another MAG has matches to Chlorobaculum, and one has matches to Candidatus Moranbacteria.
Let's classify them "officially" using sourmash and an average nucleotide identify threshold of 0.8 -
```
sourmash tax genome -g MAG*.x.gtdb.csv \
-t gtdb-rs207.taxonomy.sqldb -o MAGs \
--ani 0.8
```
This is an extremely liberal ANI threshold, incidentally; in reality you'd probably want to do something more stringent, as at least one of these is probably a new species.
This will produce a file `MAGs.classification.csv`; let's take a look:
```
cut -d, -f 1,3,5 MAGs.classifications.csv
```
Now, convert this into a lineage spreadsheet like the GTDB taxonomy, above. To do this we'll need to use a custom script [`tax-genome-to-lineage.py`](https://github.com/mblstamps/stamps2022/blob/main/kmers_and_sourmash/tax-genome-to-lineage.py) - download it and make it executable like so:
```
curl -JLO https://raw.githubusercontent.com/mblstamps/stamps2022/main/kmers_and_sourmash/tax-genome-to-lineage.py
chmod +x tax-genome-to-lineage.py
```
Now, we can run the script on the MAG classifications and produce a taxonomy file:
```
./tax-genome-to-lineage.py MAGs.classifications.csv \
MAGs.taxonomy.csv
```
Check it out!
```
cat MAGs.taxonomy.csv
```
and you should see:
```
>ident,superkingdom,phylum,class,order,family,genus,species
>MAG1_1,d__Bacteria,p__Patescibacteria,c__Paceibacteria,o__Moranbacterales,f__UBA1
568,g__JAAXTX01,s__JAAXTX01 sp013334245
>MAG2_1,d__Bacteria,p__Bacteroidota,c__Chlorobia,o__Chlorobiales,f__Chlorobiaceae,
g__Chlorobaculum,s__Chlorobaculum parvum_B
>MAG3_1,d__Bacteria,p__Bacteroidota,c__Chlorobia,o__Chlorobiales,f__Chlorobiaceae,
g__Prosthecochloris,s__Prosthecochloris vibrioformis
```
Now we can re-classify the metagenome using the combined information:
```
sourmash tax metagenome -g SRR8859675.x.gtdb+MAGS.csv \
-t gtdb-rs207.taxonomy.sqldb MAGs.taxonomy.csv | \
cut -d, -f 2,4,7 | \
sort -t, -k 3 -rn | grep order
```
Now only 56.5% remains unclassified, which is much better than before!
## Interlude: where we are and what we've done so far
::::success
To recap, we've done the following:
* analyze a metagenome's composition against 65,000 GTDB genomes, using 31-mers;
* found that a disappointingly small fraction of the metagenome can be identified this way.
* incorporated MAGs built from the metagenome into this analysis, bumping up the classification rate to ~45%;
* added taxonomic output to both sets of analyses.
::::
## How much of the metagenome is assembled??
ATLAS only bins bacterial and archaeal genomes, so we wouldn't expect much in the way of viral or eukaryotic genomes to be binned.
But... how much even _assembles_?
Let's pick a few of the matching genomes out from GTDB and evaluate how many of the k-mers from that genome match to the unassembled metagenome, and then how many of them match to the assembled contigs.
::::info
Make sure you're in the `smash` conda environment:
```
conda activate # get back to base environment
conda activate smash # activate sourmash environment
```
::::
First, download the contigs:
```
curl -JLO https://osf.io/jfuhy/download
```
this produces a file `SRR8859675_contigs.fasta`.
Sketch the contigs into a sourmash signature -
```
sourmash sketch dna SRR8859675_contigs.fasta --name-from-first
```
Now, extract one of the top gather matches to use as a query; this is "Chromatiaceae bacterium":
```
sourmash sig cat matches.zip --include GCA_016938795.1 -o GCA_016938795.sig
```
### Evaluate containment of known genomes in reads vs assembly
::::info
If you want to just start here, you can download the files needed for the below sourmash searches like so:
```
mkdir -p ~/kmers
cd ~/kmers
wget https://github.com/mblstamps/stamps2022/raw/main/kmers_and_sourmash/assembly-loss-files.zip
unzip -o assembly-loss-files.zip
```
You may also need to do
```
conda activate base
conda activate smash
```
before running sourmash.
::::
Now do a containment search of this genome against both the unassembled metagenome and the assembled (but unbinned) contigs -
```
sourmash search --containment GCA_016938795.sig \
SRR8859675*.sig* --threshold=0 --ignore-abund
```
We see:
```
similarity match
---------- -----
23.3% SRR8859675
4.7% SRR8859675_0
```
where the first match (at 23.3% containment) is to the metagenome. (You'll note this matches the % in the gather output, too.)
The second match is to the assembled contigs, and it's 4.7%. That means ~19% of the k-mers that match to this GTDB genome are present in the unassembled metagenome, but are lost during the assembly process.
Why? Any ideas?
::::success
::::spoiler Some thoughts and answers
It _could_ be that the GTDB genome is full of errors, and those errors are shared with the metagenome, and assembly is squashing those errors. Yay!
But this is extremely unlikely... This GTDB genome is entirely independent from this sample...
It's much more likely (IMO) that one of two things is happening:
(1) this sample contains _several_ strain variants of this genome, and assembly is squashing the strain variation, because that's what assembly does.
(2) this sample is at low abundance in the metagenome, and assembly can only recover parts of it.
Which do you think is more likely in this scenario?
::::
Note, you can try the above with another one of the top gather matches and you'll see it's *entirely* lost in the process of assembly -
```
sourmash sig cat matches.zip --include GCF_004138165.1 -o GCF_004138165.sig
sourmash search --containment GCF_004138165.sig \
SRR8859675*.sig* \
--ignore-abund --threshold=0
```
why, do you think?
## Summary and concluding thoughts
Above, we demonstrated a _reference-based_ analysis of shotgun metagenome data using sourmash.
We then _updated our references_ from the MAGS produced by the assembly and binning tutorial on Monday, which increased our classification rate substantially.
Last but not least, we looked at the loss of k-mer information due to metagenome assembly.
All of these results were based on 31-mer overlap and containment - k-mers FTW!
A few points:
* We would have gotten slightly different results using k=21 or k=51; more of the metagenome would have been classified with k=21, while the classification results would have been more closely specific to genomes with k=51;
* sourmash is a nice one-stop-shop tool for doing this, but you could have gotten similar results by using other tools.
* Next steps here could include mapping reads to the genomes we found, and/or doing functional analysis on the matching genes and genomes. We'll talk more about functional analysis this afternoon!
# Additional exercises and things to try!
## Visualizing `gather`-based taxonomy with metacoder
[metacoder](https://grunwaldlab.github.io/metacoder_documentation/) is a very nice R-based taxonomy viz package from the Gruenwald lab. We can run it on the command line like so:
Install a number of R packages with conda (~2 minutes) -
```
mamba install -c conda-forge -c bioconda -y r-dplyr r-metacoder r-readr
```
Download an [R script](https://github.com/mblstamps/stamps2022/blob/main/kmers_and_sourmash/sourmash_gather_to_metacoder_plot.R) and make it executable:
```
curl -JLO https://raw.githubusercontent.com/mblstamps/stamps2022/main/kmers_and_sourmash/sourmash_gather_to_metacoder_plot.R
chmod +x sourmash_gather_to_metacoder_plot.R
```
Run the R script:
```
./sourmash_gather_to_metacoder_plot.R SRR8859675.x.gtdb+MAGS.csv
```
This will produce a file [metacoder_gather.png](https://raw.githubusercontent.com/mblstamps/stamps2022/main/kmers_and_sourmash/metacoder_gather.png).
## Classify the metagenome against the entire GTDB
If you like, you can try classifying the metagenome against all 320,000 bacterial and archaeal genomes in GTDB. Start by downloading the entire GTDB reference set for k=31 from [the sourmash prepared databases page](https://sourmash.readthedocs.io/en/latest/databases.html) -
```
curl -JLO https://osf.io/k2u8s/download
```
This will create a 10 GB file, `gtdb-rs207.genomic.k31.zip`.
Now run the gather with this larger database; this will take ~32 minutes.
```
sourmash gather SRR8859675.sig gtdb-rs207.genomic.k31.zip
```
The 'gather' against the genomic representatives identified 5.3% of the metagenome; how much more do you get with the entire GTDB?
::::success
::::spoiler An answer:
```
found 23 matches total;
the recovered matches hit 5.8% of the abundance-weighted query
```
::::
## Using genome-grist to generate mapping results
(30 minutes)
[genome-grist](https://dib-lab.github.io/genome-grist/) is a workflow that automates the sourmash gather steps that we did above, and adds mapping and visualization. It's meant to eventually be a one-stop shop for metagenomes to genomes, at least for Illumina sequence.
genome-grist will do any or all of the following:
* download a metagenome from the SRA (or use a private metagenome);
* find all matching genomes in Genbank using sourmash gather;
* do taxonomic classification and provide figures;
* download all matching genomes and map the reads to them;
* build summary plots of the mapping rates;
You can install it like so:
```
pip install genome-grist
```
and now we need to configure it.
Per [the genome-grist docs](https://dib-lab.github.io/genome-grist/configuring/#preparing-information-on-local-genomes), we will configure genome-grist to make use of both GTDB genomes _and_ the MAGs we built.
First, create a single sourmash database containing the three MAGs:
```
sourmash sig cat MAG*.fasta.sig -o MAGs.db.zip
```
Next, create a subdirectory to store the actual MAG genomes (needed for mapping) -
```
mkdir MAGs
python -m genome_grist.copy_local_genomes MAG*.fasta -o MAGs/MAGs.info.csv -d MAGs.d
```
and make info CSVs for them -
```
python -m genome_grist.make_info_file MAGs/MAGs.info.csv
```
Finally, create a configuration file by executing the entire block of code below -
```shell
cat > stamps.conf <<EOF
samples:
- SRR8859675
outdir: outputs.stamps/
metagenome_trim_memory: 5e9
sourmash_databases:
- gtdb-rs207.genomic-reps.dna.k31.zip
- MAGs.db.zip
local_databases_info:
- MAGs/MAGs.info.csv
taxonomies:
- gtdb-rs207.taxonomy.sqldb
- MAGs.taxonomy.csv
EOF
```
Now we'll run genome-grist; this will take about 30 minutes to install all the software, download and prepare the metagenome, etc.
```
genome-grist run stamps.conf summarize -j 8 -p
```
## Applying spacegraphcats after genome grist
Generate sgc config file:
```
genome-grist run stamps.conf make_sgc_conf
```
then install spacegraphcats dependencies:
```
wget https://raw.githubusercontent.com/spacegraphcats/spacegraphcats/latest/environment.yml -O spacegraphcats-env.yml
mamba env create -f spacegraphcats-env.yml
```
Activate sgc environment and install sgc:
```
conda activate spacegraphcats
pip install spacegraphcats
```
Now run:
```
spacegraphcats run outputs.stamps/sgc/SRR8859675.conf search
```
## From raw metagenome reads to phyloseq taxonomy table using `sourmash gather` and `sourmash taxonomy`
Tutorial link [here](https://hackmd.io/qviirBCtT9iilWprNHjbhA?view)!
## From raw metagenome reads to taxonomic differential abundance analysis using `sourmash taxonomy` results
Tutorial link [here](https://hackmd.io/uYqDstT5RsK-OIPY1wSjhg?view)!

